Databases for Protein Structure:
[Links]
The PDB archive is the single worldwide repository of information about the 3D structures of large biological molecules, including proteins and nucleic acids. These are the molecules of life that are found in all organisms including bacteria, yeast, plants, flies, other animals, and humans.
PDBsum is a pictorial database that provides an at-a-glance overview of the contents of each 3D structure deposited in the Protein Data Bank (PDB).
It shows the molecule(s) that make up the structure (ie protein chains, DNA, ligands and metal ions) and schematic diagrams of the interactions between them.
PROSITE: Database of protein domains, families and functional sites
PROSITE consists of documentation entries describing protein domains, families and functional sites as well as associated patterns and profiles to identify them.
PROSITE is complemented by ProRule, a collection of rules based on profiles and patterns, which increases the discriminatory power of profiles and patterns by providing additional information about functionally and/or structurally critical amino acids.
Prediction of transmembrane helices in proteins
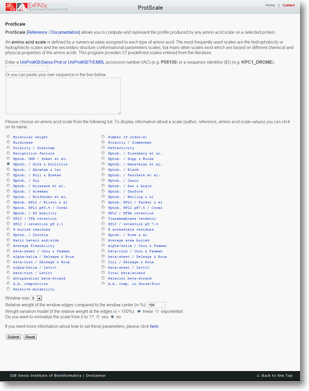
ProtScale allows you to compute and represent the profile produced by any amino acid scale on a selected protein.
An amino acid scale is defined by a numerical value assigned to each type of amino acid. The most frequently used scales are the hydrophobicity or hydrophilicity scales and the secondary structure conformational parameters scales, but many other scales exist which are based on different chemical and physical properties of the amino acids. This program provides 57 predefined scales entered from the literature.
InterPro is an integrated database of predictive protein signatures used for the classification and automatic annotation of proteins and genomes.
InterPro classifies sequences at superfamily, family and subfamily levels, predicting the occurrence of functional domains, repeats and important sites. InterPro adds in-depth annotation, including GO terms, to the protein signatures.
Pfam 26.0
(November 2011, 13672 families)
The Pfam database is a large collection of protein families, each represented by multiple sequence alignments and hidden Markov models (HMMs).
The free, collaborative 3D-encyclopedia of proteins & other molecules.